Andrew Mott at Texas A&M noticed recently that although CalcImage was correctly calculating and outputting a specified element by difference for the elemental weight percent and atomic percent .DAT files, the program was only outputting quant x-ray maps for elemental weight percents for the element by difference, but not quant x-ray maps for atomic percent element by difference.
And sure enough when we looked in the code and it's wasn't exporting the .GRD file properly. So we modified the code and now it properly outputs .GRD files for export to Surfer for elemental, atomic and oxide weight percents for the element by difference as seen here for atomic percents:
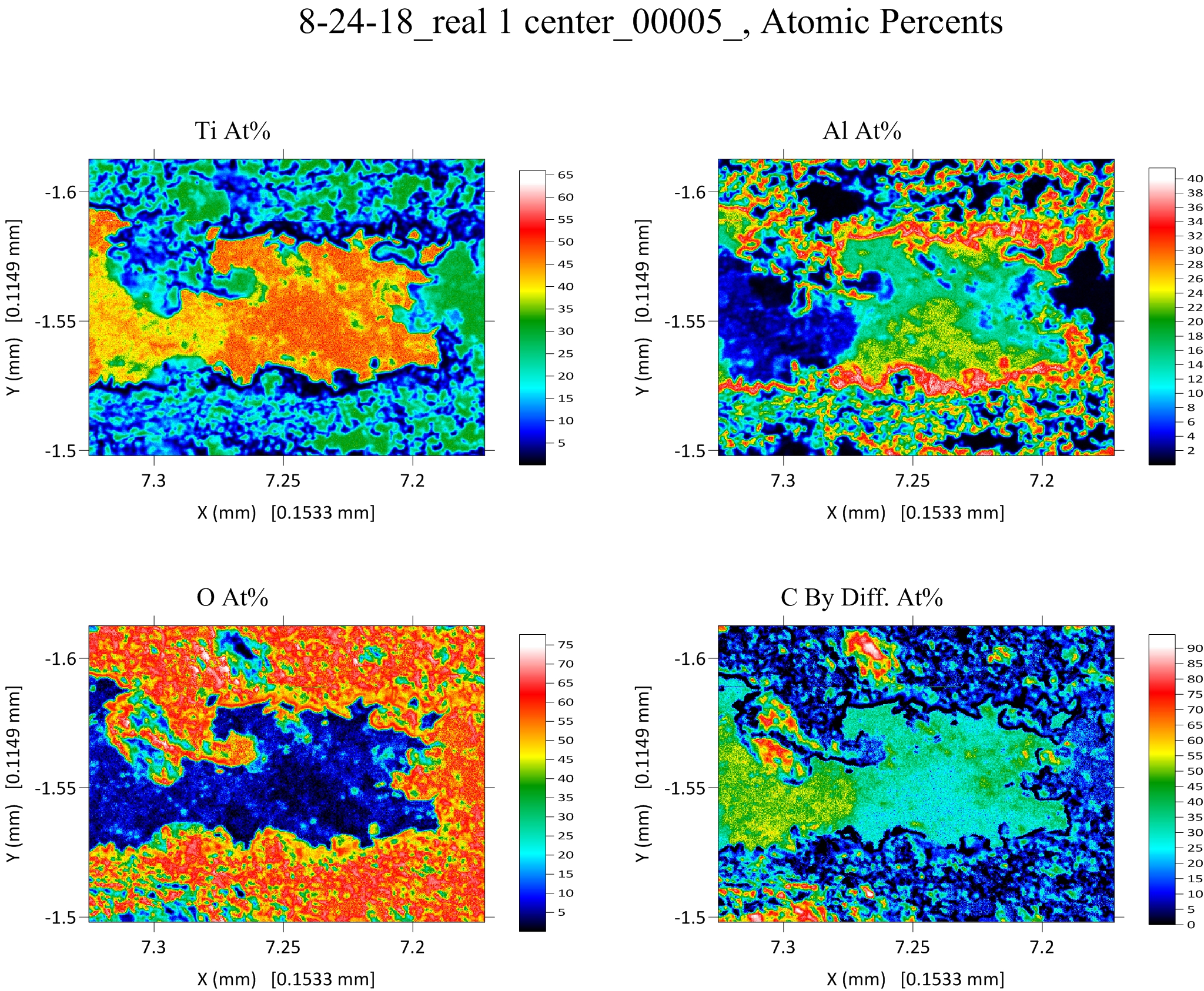
This modified code now also outputs maps for the totals, oxygen by stoichiometry, excess oxygen and mentioned already, the element specified by difference.
However, when exporting the maps to Surfer, the code automatically suppresses the total concentrations maps for atomic and element by difference output since those maps are always going to be pixels all at 100%.
One caveat: because the new code now outputs separately named .GRD files for the totals, oxygen by stoichiometry, excess oxygen and the element specified by difference elements, for both the atomic and oxide weight percent output, if you have previously calculated runs containing a totals and/or oxygen by stoichiometry quant maps and you are using atomic or oxide output, you will need to re-calculate the quant to create the newly named atomic and oxide totals and/or oxygen by stoichiometry quant maps. That is if you want to re-export the output to Surfer.
Sorry for any trouble this causes anyone. Of course, new quant map calculations in CalcImage will proceed automatically, as will any previously calculated map outputs that just used elemental weight percent output.
We think (hope) this is good for everyone but let us know what you think.